Error in Post-Training Quantization: 'Axis 2 is out of bounds for array of dimension 2' and Initializer Warnings with ONNX Model
- February 20, 2025
- 2 replies
- 969 views
Hello! I trained a neural network in MATLAB that requires an input instance of 30 features and exported it in ONNX v7 format. I am performing post-training quantization on the ST Edge AI Developer Cloud, but I am encountering issues when trying to load the dataset to check the accuracy obtained after quantization. To save this dataset in .npz format, I am using this Python script:
_____________________________________________________________
import numpy as np
import scipy.io
x_test = scipy.io.loadmat(r'C:\Users\maria\Desktop\x_test.mat')
y_test = scipy.io.loadmat(r'C:\Users\maria\Desktop\y_test.mat')
x_test = x_test['x_test']
y_test = y_test['y_test']
x_test = x_test.astype(np.float32)
np.savez("mydata.npz", x_test=x_test, y_test=y_test)
_________________________________________________________________
x_test is a matrix that contains 102 rows (number of instances) and 30 columns, while y_test is an array of 102 rows that contains the labels of the instances.
When I try to launch the quantization in the terminal, I get this error:
Executing with: {'model': '/tmp/quantization-service/3856047a-9e80-44aa-a904-d4a5d551aab9/NeuralNetwork_TreIMU_Provaaa.onnx', 'data': '/tmp/quantization-service/3856047a-9e80-44aa-a904-d4a5d551aab9/mydata.npz', 'disable_per_channel': False}
Preprocess the model to infer shapes of each tensor
axis 2 is out of bounds for array of dimension 2
2025-02-20 12:15:23.551235: I tensorflow/core/util/port.cc:113] oneDNN custom operations are on. You may see slightly different numerical results due to floating-point round-off errors from different computation orders. To turn them off, set the environment variable `TF_ENABLE_ONEDNN_OPTS=0`.
2025-02-20 12:15:23.579012: I tensorflow/core/platform/cpu_feature_guard.cc:182] This TensorFlow binary is optimized to use available CPU instructions in performance-critical operations.
To enable the following instructions: AVX2 AVX512F AVX512_VNNI FMA, in other operations, rebuild TensorFlow with the appropriate compiler flags.
2025-02-20 12:15:24.757973523 [W:onnxruntime:, graph.cc:1231 Graph] Initializer input_Scaling appears in graph inputs and will not be treated as constant value/weight. This may prevent some of the graph optimizations, like const folding. Move it out of graph inputs if there is no need to override it, by either re-generating the model with latest exporter/converter or with the tool onnxruntime/tools/python/remove_initializer_from_input.py.
2025-02-20 12:15:24.758000628 [W:onnxruntime:, graph.cc:1231 Graph] Initializer input_Shift appears in graph inputs and will not be treated as constant value/weight. This may prevent some of the graph optimizations, like const folding. Move it out of graph inputs if there is no need to override it, by either re-generating the model with latest exporter/converter or with the tool onnxruntime/tools/python/remove_initializer_from_input.py.
2025-02-20 12:15:24.758005838 [W:onnxruntime:, graph.cc:1231 Graph] Initializer fc_1_MatMul_W appears in graph inputs and will not be treated as constant value/weight. This may prevent some of the graph optimizations, like const folding. Move it out of graph inputs if there is no need to override it, by either re-generating the model with latest exporter/converter or with the tool onnxruntime/tools/python/remove_initializer_from_input.py.
2025-02-20 12:15:24.758009891 [W:onnxruntime:, graph.cc:1231 Graph] Initializer fc_1_Add_B appears in graph inputs and will not be treated as constant value/weight. This may prevent some of the graph optimizations, like const folding. Move it out of graph inputs if there is no need to override it, by either re-generating the model with latest exporter/converter or with the tool onnxruntime/tools/python/remove_initializer_from_input.py.
2025-02-20 12:15:24.758013833 [W:onnxruntime:, graph.cc:1231 Graph] Initializer fc_2_MatMul_W appears in graph inputs and will not be treated as constant value/weight. This may prevent some of the graph optimizations, like const folding. Move it out of graph inputs if there is no need to override it, by either re-generating the model with latest exporter/converter or with the tool onnxruntime/tools/python/remove_initializer_from_input.py.
2025-02-20 12:15:24.758017604 [W:onnxruntime:, graph.cc:1231 Graph] Initializer fc_2_Add_B appears in graph inputs and will not be treated as constant value/weight. This may prevent some of the graph optimizations, like const folding. Move it out of graph inputs if there is no need to override it, by either re-generating the model with latest exporter/converter or with the tool onnxruntime/tools/python/remove_initializer_from_input.py.
2025-02-20 12:15:24.758021321 [W:onnxruntime:, graph.cc:1231 Graph] Initializer fc_3_MatMul_W appears in graph inputs and will not be treated as constant value/weight. This may prevent some of the graph optimizations, like const folding. Move it out of graph inputs if there is no need to override it, by either re-generating the model with latest exporter/converter or with the tool onnxruntime/tools/python/remove_initializer_from_input.py.
2025-02-20 12:15:24.758024952 [W:onnxruntime:, graph.cc:1231 Graph] Initializer fc_3_Add_B appears in graph inputs and will not be treated as constant value/weight. This may prevent some of the graph optimizations, like const folding. Move it out of graph inputs if there is no need to override it, by either re-generating the model with latest exporter/converter or with the tool onnxruntime/tools/python/remove_initializer_from_input.py.
I don’t understand the reason for the “axis 2 is out of bounds for array of dimension 2” error because, in my network, I work with a two-dimensional matrix, and I don’t understand why it would try to access a third dimension. Additionally, I don't understand why it warns me that the initializers appear in graph inputs because I'm fairly sure they were exported as constants.
At this point, I don't know if the error is in the network or in the .npz dataset. How can I resolve this?
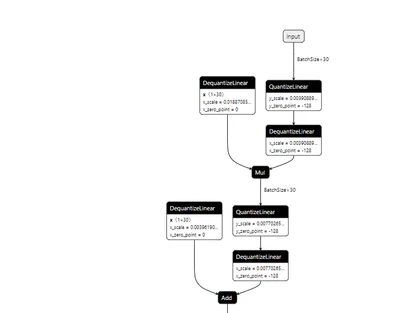
I am attaching the exported network.
Thank you in advance for your help!